Cell lines
IMR-32 cells were obtained from ATCC in March 2023. Cells were maintained in minimal essential medium, supplemented with 10% FBS, 2 mmol/L glutamine, 100 U/ml penicillin, and 100 µg/ml streptomycin, 1 mmol/L pyruvate and 0.075% NaHCO3 at 5% CO2 incubator at 37 °C. Cells were maintained in culture for up to 20 passages and tested for mycoplasma at regular intervals.
Mice protocol
Male immunocompromised NOD-SCID gamma (NOD.Cg-PrkdcSCIDII2rgtm1Wjl/SzJ) mice were purchased from The Jackson Laboratory (Wilmington, MA) between 4-6 weeks of age, and were housed five per cage with food and water ad libitum on a 12:12 light/dark cycle. Mice were given enviropak for enrichment. Mice were allowed to adjust to the new environment for two weeks before experiments were initiated. Experimental protocols were approved by the institutional animal care and use committee (IACUC) at University of Michigan, Ann Arbor, Michigan under protocol number PRO00011833. The rationale for first studying male mice was based on prior research suggesting that male mice exhibit heightened sensitivity to stress responses, particularly in models of depression and anxiety-related behaviors30. Given that our study aims to investigate the impact of food insecurity as a chronic stressor and its effects on tumor progression, we selected male mice to minimize potential variability introduced by sex differences in stress resilience and hormonal fluctuations. We fully acknowledge that sex is a critical biological variable, and future studies will incorporate both male and female cohorts to examine potential sex-specific differences in neuroblastoma adaptation to socioeconomic stressors. We have complied with all relevant ethical regulations for animal use.
Tumor inoculation and monitoring
IMR-32 cells in the exponential growth phase were prepared for injection. 1 × 105 IMR-32 cells suspended in 1:1 dilution of RPMI (Corning) and Matrigel (Corning) were percutaneously injected into the left adrenal gland of NSG mice using ultrasound guidance as previously described24. Tumors were monitored weekly for engraftment and growth using ultrasound. Tumor volumes were graphed using ½(ab2) formula. The maximum tumor size allowed by the IACUC is 2 cm in any one direction or 2000 mm³ in volume, and in none of the experiments were these limits exceeded.
Chow scarcity food insecurity modeling protocol
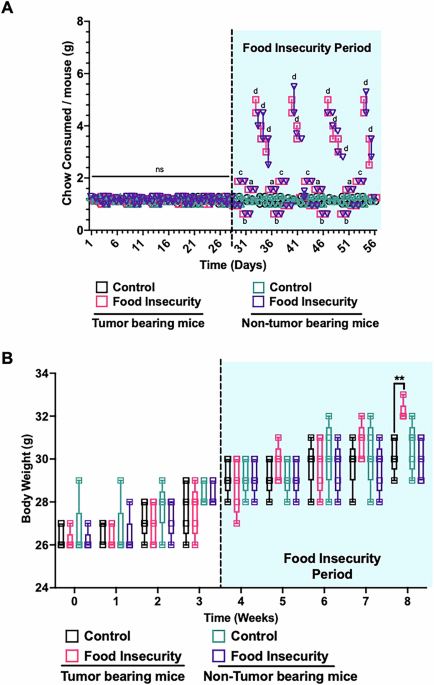
The following in vivo food insecurity protocol was adapted from Estacio et al. for neuroblastoma xenografted mice25. Following tumor engraftment verification through serial ultrasound and volume measurements, xenografted mice were randomly assigned to either the food insecurity or control groups. The food insecurity group consisted of two cages with 5 mice each, subjected to a fluctuating food availability regimen designed to simulate human food insecurity. For four consecutive weeks, the experimental groups received varied, unpredictable amounts of standard chow over four days each week, followed by three days of ad libitum access (500% representing a stress recovery period). During the four-day exposure periods, mice were provided chow at 50%, 75%, 125%, and 150% of their baseline caloric consumption, in a randomly assigned order each week (Table 1). Daily weighing of chow allowed accurate assessment of food consumption at the cage level, ensuring precise monitoring of intake patterns. This regimen aimed to emulate irregular and uncertain food access characteristic of human food insecurity, where periods of sufficient food availability are interspersed with unpredictable scarcity. The control group, also consisting of two cages with 5 mice each, was maintained on consistent, ad libitum access to standard chow throughout the study. Similar to the food insecurity group, food consumption in the control group was monitored daily. This control setup provided a stable baseline against which the effects of food insecurity on tumor progression could be measured and compared. Both control and treatment groups were handled similarly to reduce variability (Table 1).
Euthanasia and necropsy
When tumors reached 2 cm in any one direction or 2000 mm3 volume observed via ultrasound, mice were euthanized as per Unit for Laboratory Animal Medicine (ULAM) guidelines and tumors harvested. Immediately after euthanasia, tumor dimensions and weights were recorded. Portions of the harvested tumors were stored in MACS tissue freezing solution (Miltenyi Biotec) and stored at –80 °C for processing. Portions of harvested tumor were also snap frozen in liquid nitrogen and stored at -80°C for RNA and protein analysis. The remaining tumor specimen and animal carcasses were sent to the ULAM pathology core for necropsy, molecular analysis, and evaluation for metastasis.
Blood sample collection
Approximately 1% of the mouse’s body weight, roughly 150–200 µL of blood, was collected biweekly into EDTA-coated tubes from mouse submandibular veins. To keep hormonal measurements consistent, all samples were collected 3 days post-stress recovery periods at 8:00am.
Plasma hormonal quantification
All samples were harvested after three consecutive ad libitum feeding days, at the same time in the morning. Plasma was separated by centrifugation at 1500 × g for 10 min using a refrigerated centrifuge set to 4 °C. Corticosterone, adrenaline and noradrenaline, recognized as biological markers for physiological stress, were assayed from these plasma samples. Additionally, as a preliminary indicator of metastatic activity, serum levels of vascular endothelial growth factor A (VEGFA), a key promoter of angiogenesis and metastasis were quantified. All hormones were measured using commercial ELISA kits, following the manufacturer’s instructions (Abcam, CA).
Tissue hormonal measurements
For hormonal measurements from tissue, after necropsy, xenografted tumors were rinsed with ice-cold PBS (0.01 M, pH = 7.4). Tissue specimens were weighed and minced to small samples which were homogenized in PBS containing a protease inhibitor with a glass homogenizer on ice. To further separate the cells, the suspension was sonicated with an ultrasonic cell disruptor and subjected to freeze-thaw cycles. The homogenates were then centrifuged for five minutes at 5000 × g to retrieve the supernatant. Hormones were measured from obtained supernatants using ELISA kit per manufacturer’s instructions (Abcam, CA). This protocol allowed for precise quantification of catecholamines, angiogenesis marker and glucocorticoids, providing insights into the biochemical stress response within the tumor microenvironment.
Western blot
For immunoblots, 0.1 g of tumor was minced and homogenized in a RIPA buffer containing protease inhibitors and phosphatase inhibitors. Protein from tissue homogenates was measured using a BCA kit (Thermo Fisher). 25ug of protein was separated using 3–8% Tris-Acetate gels for higher molecular weight proteins and 4–20% Tris-Glycine for all other proteins (Invitrogen). For transferring, Tris-Acetate gel was incubated for 15 min in 10% methanol- containing transfer buffer before proteins were electro‐transferred to methanol-activated immobilon‐FL PVDF membranes (EMD Millipore, Billerica, MA). Membranes were then blocked with 5% milk in TBST buffer for 1 h and incubated with 1:1000 dilutions in 5% milk in TBST overnight at 4°C with primary antibodies—anti-phospho-mTOR (Cell Signaling #2974), anti-mTOR (Cell Signaling # 2983), anti‐RPTOR (Cell Signaling #2280), anti‐GBL (Cell Signaling #3274) and anti-RICTR (Cell Signaling #2114), anti-GAPDH (Cell Signaling #2118), anti-SMA Santa Cruz Biotechnology #sc-53015, anti-PARP1 (Santa Cruz Biotechnology #sc-56196), anti-MMP9 (Santa Cruz Biotechnology #sc-21733), anti-PCNA (Santa Cruz Biotechnology #sc-56). Membranes were then washed with TBST (10 min × 3), incubated with IgG HRP-conjugated secondary antibodies (Cell signaling, anti-mouse (#7076), anti-rabbit (#7074)), 1:5000 dilutions in 5% milk for 1 h at room temperature, and washed with TBST (10 min × 3). Chemiluminescence signal with ECL substrate was scanned on iBright FL1000 (Invitrogen). Signal intensities were quantified with ImageJ software (U.S. National Institutes of Health, Bethesda, Maryland, USA), and charted with Graphpad Prism version 10.0.3 (Boston, Massachusetts). All uncropped original blots are included in the manuscript under supplementary information.
PCR array
RNA was isolated from tumor tissues using Trizol and RNeasy Universal kit as per manufacturer’s instructions (Qiagen, Michigan, USA). For cDNA synthesis RT2 First strand kit was used as per manufacturer’s protocol (Qiagen, Michigan, USA). Human Neurotransmitter Receptors (GeneGlobe ID: PAHS-060Z, Cat. No.: 330231) and Human PI3K/AKT Signaling Pathway (GeneGlobe ID: PAHS-058Z, Cat. No.: 330231) PCR array were performed. Data was analyzed using Qiagen PCR analysis portal website. Two individual samples from each group were used for replicates. Genes shown in Fig. 6A, B represent those with ≥3-fold change and a p value < 0.05. Expression values were normalized to internal housekeeping controls per manufacturer’s instructions.
Immunohistochemistry
Unstained sections were cut on a rotary microtome at 4 µm thickness and mounted on glass slides. Heat-induced epitope retrieval was performed in a pH 6.2 buffer (DV2004, DIVA Decloaker, Biocare Medical) in a laboratory pressure cooker (DC2002, Decloaking ChamberTM, Biocare Medical) at temperatures alternating between 125 °C (40 seconds) and 95 °C (10 s) for a total of 60 min. Immunohistochemical staining was performed on an automated immunostainer (Biocare Intellipath, Biocare Medical). The protocol consisted of endogenous alkaline phosphatase quenching and blocking of non-specific sites (Biocare Rodent Block M, RBM961, Biocare Medical) followed by application of the primary antibody Ki67 (Cat. # ab16667, Abcam) at room temperature at the dilutions specified above. Incubation was 1 h for both primary antibodies. Detection was performed using a biotin-free polymer-based detection system (Rabbit-on-Rodent AP polymer, RMR625G, Biocare Medical) with a Fast Red chromogen (intelliPath Fast Red, IPK5017, Biocare Medical). Slides were counterstained with hematoxylin (Biocare Medical), air dried overnight, cleared in xylene, and cover slipped using a permanent mounting media. A control block with multiple types of mouse tissues was utilized as a positive control. Negative control immunostains were performed with each run using the same protocol as above with the substitution of commercially provided naïve mouse/rabbit IgG (Universal negative, IP498G20, Biocare Medical) in place of the primary antibody. Control and experimental slides were evaluated by a board-certified research veterinary pathologist (BEC) with respect to staining compartment, cell-specificity, intensity, and presence/absence of background or artifact.
Digital slide assessment and quantification
Slides were digitized on a Leica Aperio AT2 digital slide scanner (Leica Biosystems) at a resolution of 0.25 µm/pixel (20x objective). Quantitative assessment of immunohistochemical staining was performed using the open-source program QuPath v0.4.3 (github.com/qupath/qupath). Detection parameters were optimized using the positive and negative control slides. Analysis parameters for positive cell detection are given in Table 3. All result overlays were visually checked for accuracy by a board-certified veterinary pathologist (BEC).
Histology evaluation
Histological sections were evaluated using light microscopy at magnifications ranging from 20x to 600x by a board-certified veterinary pathologist. Lesions were assessed per International Harmonization of Nomenclature and Diagnostic Criteria (INHAND) consensus guidelines for rodent toxicologic pathology (available at https://www.toxpath.org/inhand.asp).
Metastatic foci characterization
Foci of metastasis were characterized as “micro” if the cells did not appear to disrupt normal alveolar architecture. The overall frequency of foci was characterized as “common” or “rare” based on the subjective ease in identifying foci. Foci were additionally characterized as small, medium, or large based on the estimated number of cells per focus (≤5, 5–10, >10, respectively).
Statistical and reproducibility
All experiments were independently repeated for a total of three trials (n = 10 per trial). Results were analyzed with ANOVA and Student’s t test using GraphPad Prism, Boston, MA, USA. All data are presented as the mean ± the standard deviation (SD) of three experiments, ns p > 0.05, a,*p < 0.05, b,**p < 0.01, c,***p < 0.001, d,****p < 0.0001.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.