Study design and participants
This study employed a retrospective cohort study design and initially included data from 26,540 pregnancy-related records at the Second Affiliated Hospital of Shaanxi University of Chinese Medicine, encompassing deliveries and cases of threatened abortion, from women aged 20 to 40 years old between January 5, 2022, and March 29, 2024. After applying exclusion criteria, the final number of participants was 3,253. Sample size was determined based on the available retrospective data from eligible patients during the study period .The study protocol was reviewed and approved by the Ethical Committee of the Second Affiliated Hospital of Shaanxi University of Chinese Medicine (Approval Number: LW2024004-1, Date: 2024/4/23), in accordance with the ethical guidelines of the latest version of the Declaration of Helsinki. Given the retrospective nature of this analysis, all patient information was anonymized, and individual participant informed consent was not required.
This research complies with both the Strengthening the Reporting of Observational Studies in Epidemiology (STROBE) guidelines for observational studies and the Transparent Reporting of a Multivariable Prediction Model for Individual Prognosis or Diagnosis (TRIPOD) guidelines for prediction model studies. The STROBE checklist is included in Supplementary File 1, and the TRIPOD checklist is included in Supplementary File 2.
During the data screening phase, the research team adopted a rigorous and well-organized step-by-step strategy to ensure data quality: first, the algorithm was used to batch screen out patient data containing a large number of missing values and zero values; then medical experts conducted manual verification of values that were significantly deviated from the normal range to further remove abnormal data. For the processing of patient data types, if the same patient has both negative (health examination) and positive (threatened abortion) data, only the positive first visit data were retained; if the patient has only a single type of data, the first visit data will be retained, and for patients with threatened abortion, the data must be strictly limited to examination records within the first three months of pregnancy. In addition, the study systematically excluded outlier data, as well as multiple pregnancy cases with at least three births, patients diagnosed with missed abortion, damaged eggs, and planned termination of pregnancy. Patients with major diseases such as severe coronary artery disease, stroke, malignant diseases, and lost follow-up were also excluded. Ultimately, 1,764 pregnant women with threatened abortion and 1,489 healthy pregnant women without abortion were included. The inclusion criteria were: (i) diagnosis with intrauterine pregnancy was based on clinical ultrasonography; (ii) the threatened abortion group included women with pregnancy-related vaginal bleeding, while the normal pregnancy group included women without such bleeding; (iii) aged between twenty and forty years old. The exclusion criteria were as follows: (i) multiple pregnancies with at least three births; (ii) diagnosed with missed abortion egg damage or planned termination of pregnancy; (iii) serious medical conditions such as severe coronary artery disease, stroke, or malignant diseases; (iv) lost to follow-up.
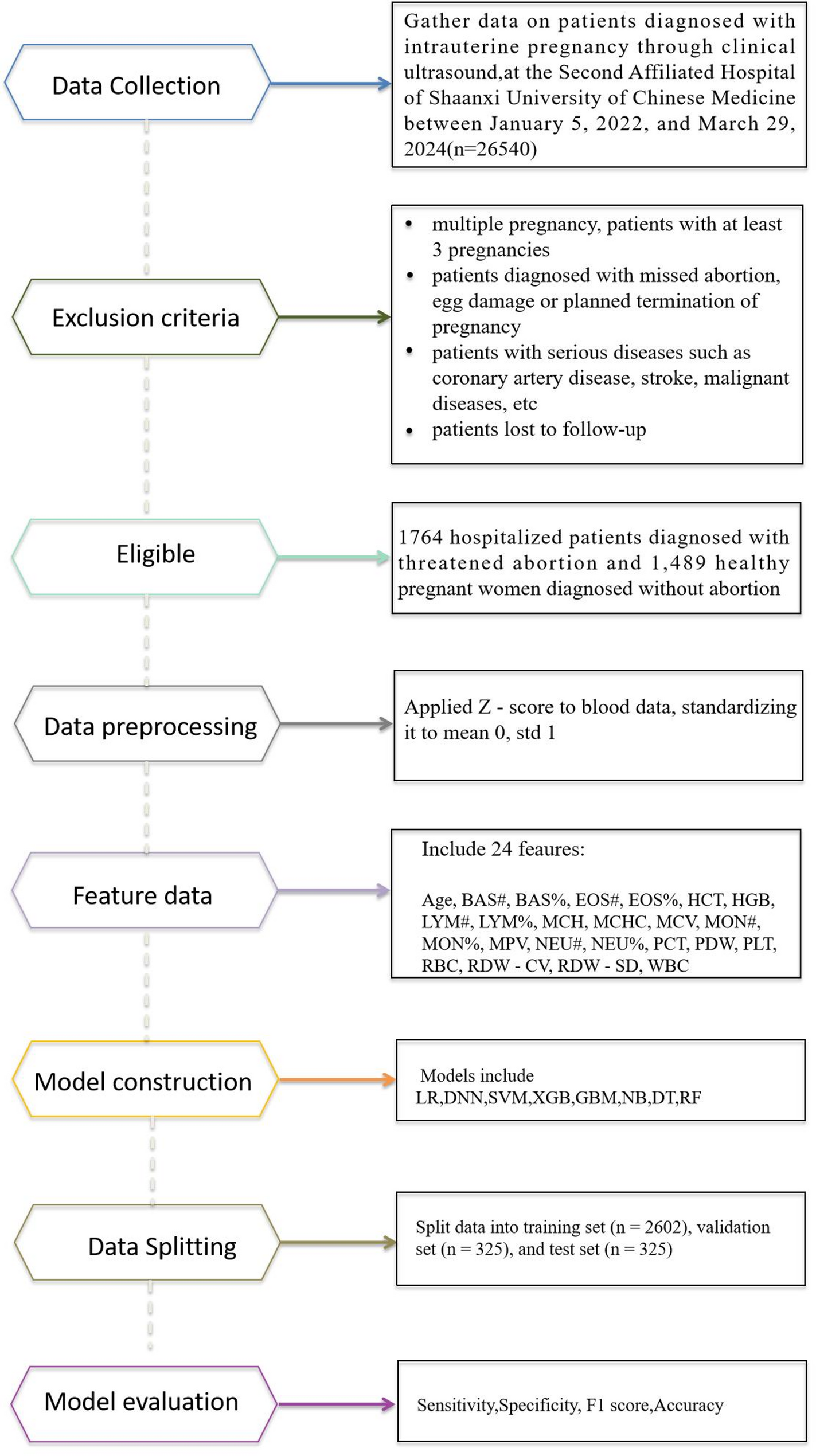
Based on the commonly – adopted ratio of training set : validation set : test set = 8:1:1, the screening data was partitioned. Specifically, 2602 cases were allocated to the training set, 325 cases to the validation set, and 325 cases to the test set. After data standardization, eight ML algorithms were used in this study to adapt the experiment and produce the final results (Fig. 1).
Flowchart of the ML development model for predicting threatened abortion
Data collection and preprocessing
We meticulously gathered blood test data from all participants to create a dataset for analysis. Upon admission, routine blood tests were performed on samples collected from each patient. Various advanced analyzers were utilized, including the Mindray Automatic Blood Cell Analyzer BC-7500, Abbott CELL-DYN series blood cell analyzer, Sysmex Europe-XN-1500 and XN-2800 automatic blood cell analyzers, and the Mindray BC-6000-Plus automatic blood cell analyzer.
In order to eliminate the interference of dimensional differences and data fluctuations among blood routine indicators on the analysis results, improve data comparability and strengthen the stability of model training, this study uses the Z score normalization method to standardize the original data. It should be noted that the Z score normalization operation and the SPSS-based statistical analysis both act directly on the original data set, and there is no fixed order of execution between the two. This method maps the original data to a standard normal distribution with a mean of 0 and a standard deviation of 1 through mathematical transformation. The specific conversion formula is as follows:
(z=frac{{x – mu }}{sigma })
In this equation, Z denotes the normalized value, X is the raw data point, µ is the mean of the dataset, and σ is the standard deviation. This normalization step allows for a fair comparison across different data points and bolsters the model’s resilience to data variability.
Model construction and optimization
To construct predictive models for early screening of threatened abortion, we employed eight ML models, each selected for its unique strengths and suitability for addressing specific predictive challenges. These models include:
LR: a widely used model for simple binary classification tasks, particularly effective in scenarios where interpretability is crucial. LR has been extensively applied in fields such as chronic kidney disease and heart disease, making it a reliable choice for our study [24,25,26].
DNN: known for its ability to capture complex patterns in data through multiple layers of interconnected nodes. DNNs excel in identifying intricate relationships within large datasets, making them highly effective for tasks requiring nuanced understanding [27, 28].
SVM: a robust model designed for classification in high-dimensional spaces. SVMs are particularly adept at finding the optimal hyperplane that separates different classes, making them ideal for datasets with many features [29]. The SVM model employed an radial basis function (RBF) kernel function to map data to a high-dimensional space and handle nonlinear relationships Therefore, in this experiment, the SVM model uses the RBF kernel function.
XGB: an advanced tree-based learning algorithm that combines multiple weak learners to form a strong predictive model. XGB is known for its efficiency and performance, offering high accuracy and scalability [30].
GBM: a models that enhance predictive accuracy through sequential learning, where each new model corrects the errors of the previous ones. GBMs are particularly effective in improving model performance iteratively [31].
DT: The decision tree model is a supervised learning algorithm based on a tree structure for classification or regression. It recursively divides data into different subsets to form a series of decision rules to achieve efficient prediction and classification of new samples [32].
NB: a straightforward and effective model, especially for text classification tasks. NB relies on Bayes’ theorem and assumes independence between predictors, making it computationally efficient and easy to interpret [33].
RF: an ensemble learning method that builds multiple decision trees and merges their predictions to produce a more accurate and stable output. RFs are highly robust, particularly when dealing with large and complex datasets [34].
Python (version 3.7.0) was used to ensure that the training process of all models followed a unified standard. The data was divided into training, validation, and test sets in a ratio of 8:1:1. We standardized the training set using the fit_transform function and applied the same transformation to the validation and test sets using the transform function.
To ensure the robustness and generalization of the constructed ML models, we used RandomizedSearchCV for hyperparameter optimization. This method is more efficient than grid search, especially in high – dimensional parameter spaces, as it reduces computational demands while seeking optimal configurations. Based on Hyperparameter Configurations and Search Space and considering the model’s complexity, we designed a search space with multiple configurations. The hyperparameter scope included an exponentially decaying learning_rate to adapt to different training phases, varying hidden_layer_sizes to explore network depth and width, testing ReLU, tanh, and sigmoid activation functions for non – linear expressiveness, and setting ‘class_weight=”balanced”’ to counter sample imbalance. The optimization and evaluation were done via five – fold cross – validation with Adam and SGD algorithms [35].
Model evaluation
To evaluate model performance and prevent overfitting, we used tenfold cross-validation. The performance metrics included true positive (TP), true negative (TN), false positive (FP), and false negative (FN). Sensitivity, specificity, F1 score, and accuracy were calculated to provide a comprehensive evaluation of the models. The calculation formula for the evaluation index is as follows:
Sensitivity = TP/(TP + FN).
Specificity = TN/(TN + FP).
F1 score = (2·precision·recall)/(precision + recall).
Accuracy = (TP + TN)/(TP + FN + TN + FP).
SHAP model explained
SHAP, based on Shapley values, is a post-hoc interpretation tool that quantifies feature contributions to predictions, ensuring local accuracy, handling missing data, and maintaining consistency. Introduced by Lundberg et al. [36], SHAP identifies influential features by calculating their impact on predictive power, enhancing model accuracy and interpretability. The model’s output is the sum of individual contributions from all features.
(fleft( x right)={phi _0}+sumlimits_{{i=1}}^{N} {{phi _i} cdot {x_i}} )
Here, (:{{varnothing}}_{0}:)is the baseline prediction, N is the number of features, (:{text{x}}_{text{i}}:)is the value of the i-th feature, and (:{{varnothing}}_{text{i}}) is the SHAP value for the i-th feature, which signifies its effect on the model’s output. The SHAP value is derived from a weighted average of the model’s predictions across all possible combinations of features, with the weights based on the subset size and feature order. The SHAP is given by:
({phi _i}left( f right)=sumnolimits_{{S subseteq Nbackslash left{ i right}}} {frac{{left| S right|!(left| N right| – left| S right| – 1)!}}{{left| N right|!}}} left[ {f(S cup left{ i right} – f(S)} right])
The Shapley value for the i-th feature is computed as a weighted average that considers all possible subsets of features, with the weights assigned based on the subset’s size and the feature’s position within it. Specifically, f(S∪{i}) denotes the model’s prediction when the features within subset S as well as the i-th feature are included, while f(S) corresponds to the model’s prediction with only the features in subset S considered. This distinction allows for an assessment of the i-th feature’s individual contribution to the model’s overall prediction.
Statistical analysis
Data analysis was performed using IBM SPSS Statistics 26. Research data, including the basic information and routine blood test results of threatened abortion patients, were retrieved from the hospital information system. In constructing the dataset, patients with threatened abortion were designated as the positive sample group, and healthy pregnant women were set as the negative sample group, resulting in a dataset comprising 24 attributes. Given that all 24 characteristic attributes are quantitative variables, the following statistical strategy was adopted to compare the hematological parameter differences between pregnant women with threatened abortion and healthy pregnant women: For features that conform to a normal distribution or are skewed but meet the condition of “standard deviation < mean/3,” independent sample t-tests were conducted. Prior to performing the t-tests, Levene’s test for equality of variances was used to assess the homogeneity of variances between groups. If the p-value from Levene’s test was greater than 0.05, indicating equal variances, the standard Student’s t-test was applied; if the p-value was 0.05 or less, indicating unequal variances, Welch’s t-test was used instead. The statistical results were presented as mean ± standard deviation (mean ± SD). For features that neither follow a normal distribution nor meet the aforementioned skewed distribution conditions, the Mann-Whitney U test was applied, with results presented as median (upper quartile, lower quartile). Additionally, the Bootstrap method was utilized to estimate confidence intervals, thereby enhancing the reliability of the statistical findings.