- Public holiday declared in Sindh on March 11 for Youm-e-Ali samaa tv
- Three-day holiday announced for Islamabad schools from March 24 Geo News
- Federal govt announces spring holidays for Islamabad schools The Express Tribune
- Sindh educational…
Author: admin
-

Public holiday declared in Sindh on March 11 for Youm-e-Ali – samaa tv
-
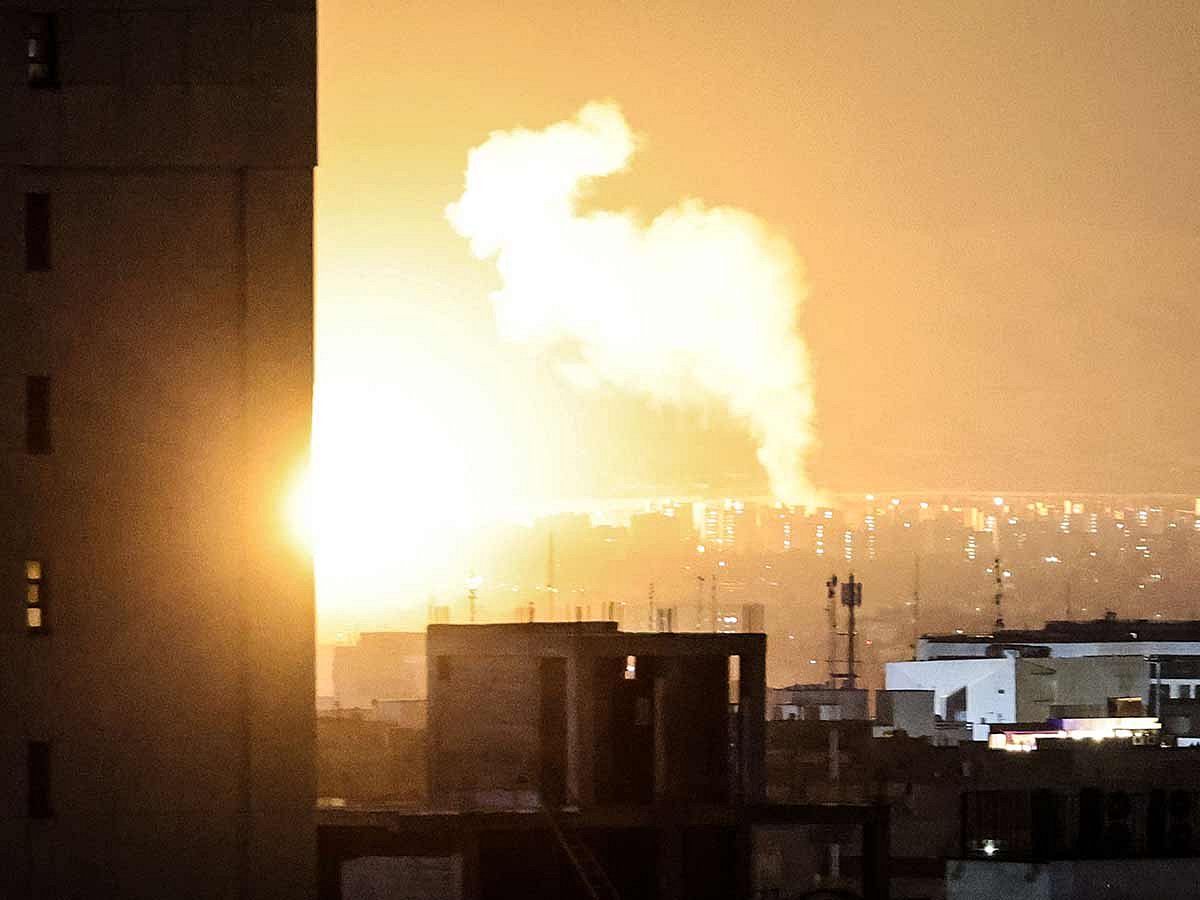

Limited GCC flights; warning over sensitive videos
The UAE air defences intercepted missiles and drones from Iran, with authorities fully prepared to counter threats and safeguard national security. Airports and airlines have resumed limited flights, while early spring breaks have been announced…
Continue Reading
-
Adults’ exposure to diverse microbes may worsen allergic conditions
The “hygiene hypothesis” suggests exposure to diverse types of microbes may protect against developing diseases caused by allergens, but a new Cornell University study in mice reveals that adults’ exposure to diverse microbes and…
Continue Reading
-

Directors’ Deals: Plus500 trio ships £67.2mn in shares to Goldman Sachs – Financial Times
- Directors’ Deals: Plus500 trio ships £67.2mn in shares to Goldman Sachs Financial Times
- Plus500 regulation, tech heads sell GBP1.8 million in shares marketscreener.com
- Plus500 Discloses Series of Insider Share Sales by Chief Regulation Officer TipRanks
- Plus500 CTO Discloses Sale of 20,000 Shares Under UK MAR Rules TipRanks
Continue Reading
-
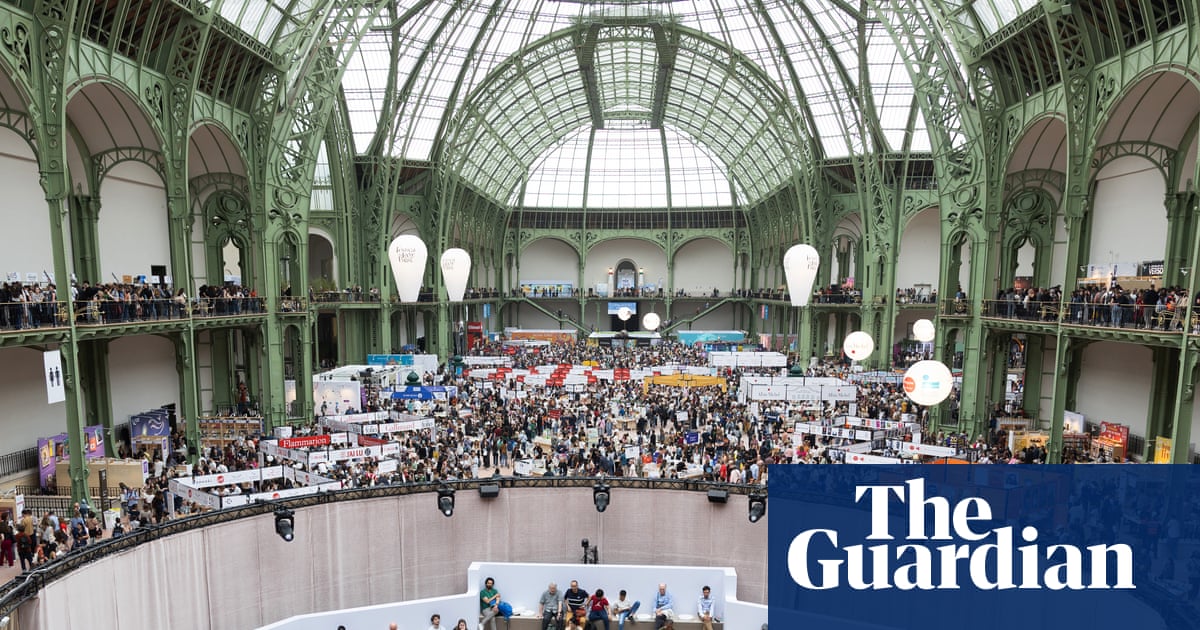

Amazon pulls sponsorship from Paris book festival after booksellers’ association boycott | Books
Amazon has withdrawn from the Paris book festival after a boycott by France’s booksellers’ association prompted a row over the company’s sponsorship of the event.
The festival, due to take place from 17 to 19 April, will now go ahead without…
Continue Reading
-

Russia is helping Iran to target US military assets in Middle East
Unlock the White House Watch newsletter for free
Your guide to what Trump’s second term means for Washington, business and the world
Russia has shared targeting intelligence with Iran, including locations of American military assets in the Middle…
Continue Reading
-
Exclusive | Kuwait Cuts Oil Production as Fallout From Iran Conflict Intensifies – WSJ
- Exclusive | Kuwait Cuts Oil Production as Fallout From Iran Conflict Intensifies WSJ
- Kuwait Cuts Oil Production as Storage Fills Up WSJ
- Iraqi Supply Loss Could Expose the Real Limits of OPEC Spare Capacity Crude Oil Prices Today | OilPrice.com
Continue Reading
