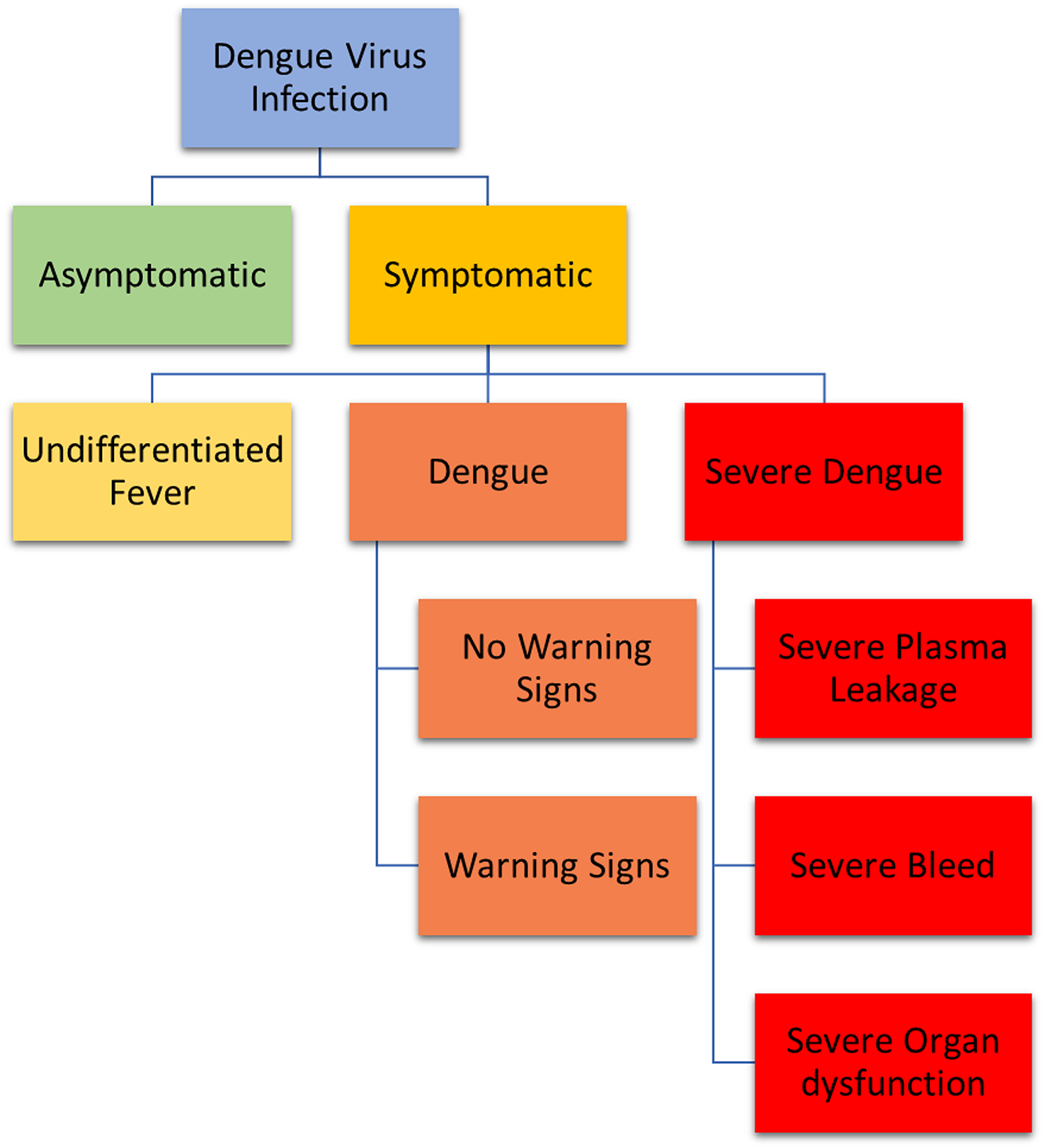
Dengue infection is one of the most prevalent communicable diseases mainly caused by the widespread thriving of the Aedes aegypti and Aedes albopictus mosquitoes. In 2022, approximately 3,766,153 cases were reported worldwide [1]. Dengue infection is caused by the transmission of one of the four dengue serotypes (DENV1, DENV2, DENV3, and DENV4) to humans via Aedes aegypti as the primary vector and Aedes albopictus as a secondary or potential vector. The clinical management of dengue infection is complex as the symptoms and manifestations of the infection is a continuum that range from asymptomatic to severe illness, which may also lead to death [2]. To enable rapid and appropriate clinical management, the World Health Organization [3] (WHO) has established a classification scheme for dengue cases, which can be categorized into dengue without warning signs, dengue with warning signs, and severe dengue (Fig. 1). While most cases are self-healing, a notable minority can progress to severe dengue, which is characterised by plasma leakage, bleeding, organ dysfunction, or potentially fatal outcomes. Recognition of severe dengue is a major challenge in the clinical setting and often requires the manifestation of the symptoms of dengue shock syndrome for accurate recognition.
Dengue cases classification and level of severity by WHO [3]
Current treatment options for dengue infections encounter numerous limitations and challenges, primarily due to the lack of specific antiviral drugs, complicated by the diverse nature of the virus, which comprises four distinct serotypes. To date, there are no licensed therapeutic agents against the dengue virus. Existing approaches are largely focused on providing supportive care, prioritizing symptom alleviation rather than directly targeting the dengue virus. Numerous clinical trials have been conducted to explore therapeutic interventions for acute dengue fever, and several more are currently underway (Table 1). These interventions target the virus, host’s immune responses, or host factors necessary for the dengue virus replication cycle. In some clinical studies, patients with severe forms of dengue fever, such as dengue haemorrhagic fever (DHF) or dengue shock syndrome (DSS), exhibited more than ten-fold higher viraemia in the early stages of infection than patients with non-severe dengue fever. This observed pattern has led to the hypothesis that the administration of antiviral therapy during the early phase of dengue infection may help to reduce the severity of clinically apparent disease by lowering viremia levels [15,16,17].
To date, there are several antibodies available against the dengue virus that target the lateral ridge or A-strand of the envelope protein (E), particularly the third domain (EDIII). These antibodies exhibit strong neutralizing capabilities against the majority of all four serotypes. However, antibodies also have disadvantages, such as high production cost, large batch-to-batch variation, high immunogenicity and potential to induce antibody-dependent enhancement (ADE) that may exacerbate the degree of cross-serotype infection. ADE in dengue virus infection refers to a process where antibodies, due to the low binding affinity unable to neutralize the virus instead facilitates a more efficient infection by enhancing fusion, while bypassing early antiviral responses, particularly in primary human macrophages. This mechanism helps explain why secondary dengue infections can be more severe and is a key concern in dengue vaccine development [18, 19]. One class of molecular recognition elements (MREs) that have advantages that recompense for these issues associated with antibodies are aptamers. Hitherto, aptamers have been widely utilized in many applications including diagnostics and therapeutics [20]. The enormous potential of aptamers is also equally capitalized for dengue therapeutics. In this review, the use of aptamers as an antiviral and therapeutic agent against dengue infection is emphasized.
Aptamers as antiviral agents
Aptamers are single-stranded DNA or RNA molecules that can fold into complex tertiary structures, allowing them to bind specifically to their cognate target molecule. As they can mimic the interaction between antibodies and antigens, aptamers are often referred to as synthetic antibodies [21]. Due to their much smaller size, aptamers can effectively reach targets that were previously inaccessible by antibodies or located intracellularly. Aptamers also have reduced immunogenicity due to their much smaller size as compared to antibodies [22].
Aptamers have several appealing properties compared to antibodies, making them a promising alternative for antibodies in various applications, including therapeutics. Aptamers can be conveniently manufactured through chemical synthesis with reduced cost and production time while demonstrating no batch-to-batch variations [23]. Unlike antibodies, which are irreversibly denatured at room temperatures or higher, aptamers can withstand ambient and elevated temperatures, reverting to their original conformations when the optimal temperature is restored. In addition, aptamers have dissociation constants that can reach as low as the picomolar range [24]. Aptamers can also be conveniently labelled and modified with a range of reporter molecules, linkers, and additional functional groups [25, 26].
Aptamers have demonstrated successful applications in therapeutics and diagnostics for various pathogens, including parasites, bacteria, and viruses [27, 28]. Due to their specific binding to target proteins, they often inhibit protein-protein interactions and can thereby induce therapeutic effects such as antagonism (Fig. 2). To date, numerous aptamers have been developed against many viruses, for example against severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2), foot-and-mouth disease virus (FMDV), human immunodeficiency virus-1 (HIV-1) and influenza A viruses [29,30,31,32].
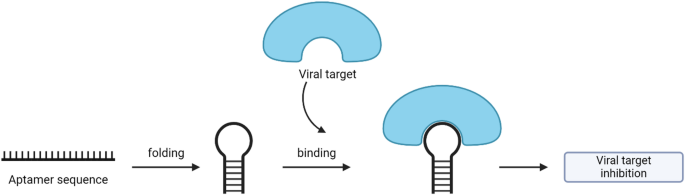
A schematic representation of aptamer binding towards a target (Created with BioRender.com)
The first therapeutic aptamer, Macugen (pegaptanib sodium), was approved by the United States Food and Drug Administration (FDA) in 2004 for the treatment of Age-related Macular Degeneration (AMD) [33]. Macugen is a 28-base ribonucleic acid aptamer that inhibits vascular endothelial growth factor (VEGF), a protein that promotes the growth of new blood vessels and often makes the blood vessels leaky. A number of clinical trials are currently underway, in which aptamers are being used for various therapeutic purposes.
Aptamers: basic and SELEX process
Conventionally, aptamers are produced by a technique known as SELEX (Systematic Evolution of Ligands by Exponential Enrichment), which involves three main steps: (a) positive selection by incubation of a randomized oligonucleotide library with the target, (b) separation of the bound nucleic acid ligands from the unbound ones, and (c) subsequent amplification of the specific target-binding candidates and their conversion into single-stranded forms (Fig. 3) [24]. PCR amplification of the specific candidates is made possible by the presence of the constant regions that flank the randomized region, which serve as the primer binding sites [34].
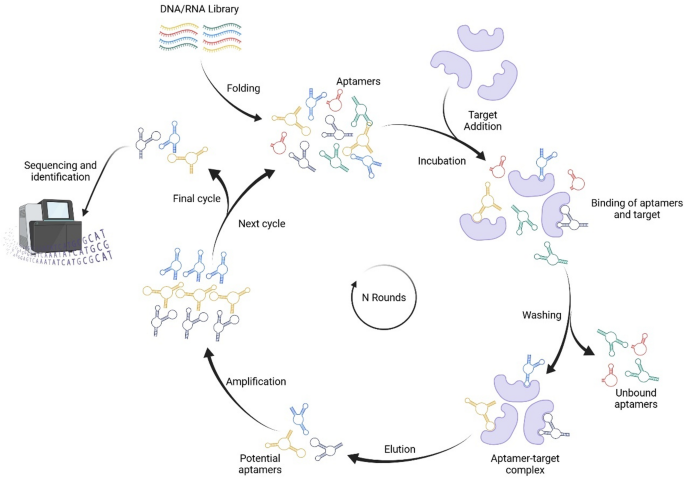
A schematic representation of conventional Systematic Evolution of Ligands by Exponential Enrichment (SELEX) procedure (Created with BioRender.com)
Counter-selection is also performed prior to positive selection by incubating the nucleic acid pool against the non-target molecules. This step is essential in discarding the non-specific binders from the library [35]. Typically, these SELEX steps are repeated from 8 to 15 times to deconvolute the nucleic acid pool, which may contain target-specific binders with increased binding affinity and specificity. This nucleic acid pool is then cloned and sequenced [36]. Following the sequence analysis, sequence classes with the highest frequency of appearance are identified as potential aptamers.
To further increase their binding affinity or specificity, the isolated aptamers can also be rationally modified. Truncation, for example, is a process that often utilizes computational methods to remove the nucleotides that are not directly involved in the binding of the aptamers to the target molecules, which are commonly located in the stem region [37]. In most cases, the active interaction sites of the aptamers are usually located in the loop region; thus they should be retained throughout the modification [38]. Secondary structural prediction tools such as mFold and RNAStructure webserver are commonly used to predict the secondary structures of the truncated aptamers [37, 39]. In addition, truncated aptamers have been shown to have higher stability and binding affinity towards the target molecules as compared to the full-length counterparts [38, 40,41,42].
Aptamers against dengue virus
The use of aptamers in targeting specific dengue viral components holds immense potential in revolutionizing the landscape of dengue viral research and management. Several aptamers have already been developed against the multiple proteins present in the virus and used as diagnostic tools for the detection of dengue virus [43]. Aptamers are also utilized as potential antiviral agents against dengue viruses. These aptamers target viral proteins (capsid, envelope domain 3) and enzymes (methyltransferase) of DENV2.
Whole virus
Amano et al. [44] employed the DENV-1 virus-like particles (VLPs), a non-infectious antigenic determinants-presenting form of the native virus, for the selection of anti-DENV aptamers through 10 rounds of SELEX. Surface plasmon resonance (SPR) analysis then revealed that the most frequently occurring sequence, named DENV-Apt, exhibited high affinity towards the DENV-1-VLPs. Truncation was then performed on this candidate, yielding a 41-mer candidate, named DENV-Apt_41, which demonstrated improved binding affinity towards the VLPs, as compared to its parent aptamer. Despite this, the truncated variant showed no promising ability in blocking the membrane fusion of the virus on HEK293T cells, indicated by the persistent luciferase signals following the challenge. As the DENV-Apt_41 does not able to block the fusion of the virus while binds to all the four serotypes, this aptamer can serve as a promising delivery tool of siRNA.
Moreover, it also displayed potential therapeutic activity against DENV-2 in vitro challenge. The application of DENV-Apt_41 reduced the viral load approximately by 50% on Day 3 of inoculation, confirmed through PCR analysis. Regardless of the therapeutic ability of this aptamer in reducing the viral replication, possibly through mechanisms apart from the membrane fusion, this study focused mainly on the application of this aptamer for the targeted delivery of silencing RNA (siRNA) to inhibit the viral replication [44].
Capsid (C) protein
The capsid (C) protein functions as a structural component of the DENV virion, encapsulating its single-stranded RNA (ssRNA) genome. It consists of four alpha helices with an unstructured amino terminus and is arranged in parallel homodimers. One side of the capsid dimer exhibits a significant electrostatic interaction, potentially playing a role in RNA binding. In contrast, the opposing side of the capsid dimer has a hydrophobic cleft, suggesting possible interactions with lipids [45, 46]. The capsid protein not only maintains the structural integrity of the viral particle but also plays a crucial role in viral replication and host-cell interaction [47].
During viral replication, the capsid protein assists in the formation of new viral particles by packaging the replicated RNA genome into new virions. Additionally, it interacts with host cell components during viral entry and uncoating, facilitating the release of the viral genome into the host cell cytoplasm, a critical step in initiating viral replication. While the integrity of the C protein does not depend on its amino terminus, deletions in this region can lead to defects in particle formation, as demonstrated previously in human cells [48, 49].
Balinsky et al. [50] investigated the interaction of the DENV C protein with the multifunctional host protein nucleolin (NCL). NCL is mainly present in the nucleolus [51] but can also be found in the cytoplasm and on the plasma membrane [52]. NCL plays roles in various cellular functions, including ribosome formation, protein transportation, chromatin modification, translation control, and maintenance of RNA processing and stability [51]. One aptamer specific against NCL termed AS1411 aptamer, is an antiproliferative G-rich sequence that is currently in Phase II clinical trials as a potential anticancer agent [53]. Although originally developed for cancer therapy, AS1411 shows potential as an antiviral agent by disrupting the interaction between nucleolin and DENV capsid protein critical for DENV replication. Pharmacokinetically, AS1411 is typically administered intravenously due to poor oral bioavailability and demonstrates rapid systemic distribution. It is stable in circulation owing to its G-quadruplex structure, which provides resistance against nuclease degradation. AS1411 is primarily cleared through renal excretion and exhibits a biphasic elimination profile with a plasma half-life of approximately 5 to 10 h. While direct pharmacokinetic studies in the context of dengue are limited, AS1411’s selective targeting, cellular uptake, and stability support its potential application as an antiviral agent against DENV. Since C protein interacts with NCL to play a crucial role in DENV replication by promoting virion morphogenesis and the production of infectious DENV progeny, AS1411 aptamer can be used as a potential dengue therapeutic agent by blocking the interaction between these two proteins. To further determine whether AS1411-mediated interference in DENV C-NCL interactions can potentially reduce DENV replication, treatment with various concentrations of AS1411 was performed. The treatment with AS1411 showed a dose-dependent reduction in DENV titer, with the maximum reduction occurring at a concentration of 5 µM [50]. Moreover, the inhibition of DENV C-NCL interaction by AS1411 aptamer led to a significant reduction in the viral titers following DENV infection. This strongly suggests that the AS1411 aptamer can be a potential dengue therapeutic agent by antagonizing the interaction between the DENV C protein and NCL.
Envelope domain III (E3) protein
The envelope protein (E) is a significant antigenic feature of DENV. It plays a crucial role in recognizing host cell receptors and facilitating the fusion of the virion with the endosomal membrane during cell entry. The envelope is organized into three domains: the central domain I (E1), the dimerized domain II (E2), and the IgC-like domain III (E3). The mechanism of the fusion involving both E2 and E3 leads to the internalization of DENV, as proposed by Modis et al. [54]. Under acidic conditions in the endosome, the E2 protein undergoes an upward flipping, revealing its fusion loop, which inserts into the endosomal membrane. This initiates trimerization, leading to the formation of a prefusion intermediate. This trimerization process extends from the fusion tip of the E protein downward to E1 at the base of the trimer. Next, the E3 protein undergoes rotation and shifting, displacing the trimers. As a result, the endosomal and viral membranes fuse. This membrane fusion may require coordinated twisting actions involving a subset of trimeric E proteins.
E3 has been targeted for the development of a thioaptamer, DENTA-1 [55]. A thioaptamer is a modified form of an aptamer in which one or more of the non-bridging oxygen atoms in the phosphate backbone of nucleotides are replaced with sulfur atoms. This creates a phosphorothioate linkage instead of the normal phosphate linkage found in DNA or RNA. This modification confers enhanced stability, specificity and affinity to thioaptamer as compared to the unmodified aptamers. A total of 38 potential thioaptamer candidates were identified after 5 cycles of SELEX. Four dominant thioaptamers, named DENTA-1 to DENTA-4, were subjected to filter binding assay for the validation of thioaptamers/E3 proteins complex formation. DENTA-1 exhibited the best binding affinity with a Kd value of 154 ± 40 nM. The interaction of the E3 protein with the aptamer was predicted to take place at H316, G317, T318 and I319. These residues are also targeted by the IgG2a neutralizing monoclonal antibody 1A1D-2, which are specific against the E3 protein [56]. DENTA-1 neutralizes the virus through its binding to E3 by distorting its conformations, which are required for membrane fusion or the initiation of pathogenesis. Due to their neutralization capacity, DENTA-1 can also be a potential therapeutic agent of dengue infection.
S15 is another example of a DNA aptamer with therapeutic capacity against E3 of DENV. Chen et al. [43] performed 15 rounds of SELEX with an initial library size containing approximately 6 × 104 oligonucleotides. Following sequencing, twelve different potential aptamer candidates were identified, with S22 and S15 being the most dominant aptamer. These candidates were then verified by fluorescence quenching experiment. It was revealed that the fluorescence emission is directly reduced by the formation of the E3/aptamer complex. One candidate, S15 reported the lowest dissociation constant (Kd) value of 200 nM, which is lower than the Kd value of 500 nM exhibited by the S22 aptamer. Therefore, S15 was selected as the best candidate and used for the subsequent assays. Epitope mapping revealed significant conformational perturbation of at least five residues (Q316, H317, G318, T319 and I320) in the conserved loop region of E3. Intentional mutations performed on certain residues (Q316G, I320A, G318P) resulted in higher Kd values of the S15, suggesting that these residues act as the main interaction sites for S15 binding. In addition, the plaque reduction neutralising test (PRNT) showed that S15 is capable of completely neutralizing DENV-2 NGC at the concentration of 100 µM. For DENV-1 HAWAII, DENV-2 NGC, DENV-3 H-87 and DENV-4 H241 treated with S15, the IC50 values obtained were 1.7 µM, 4.2 µM, 5.7 µM and 5.8 µM, respectively. To determine the binding sites of the aptamer, the authors performed some modifications to the actual S15. First, S15G3 was generated following the modifications on the G-rich regions of the aptamers by removing nucleotides at both 5’ and 3’ ends, resulting in the increased Kd values ranging from 1.1 to 3.46 µM, indicating that these terminal nucleotides are essential for the binding of S15 to the E3 protein.
Methyltransferase
Methyltransferase (MTase) is an enzyme found in flavivirus responsible for the methylation of the N-7 position of the guanine cap and the 2’-OH position of the first nucleotide of the viral genome. The MTase of flaviviruses is highly specific as it only recognises and methylates their viral RNA, particularly at the 5’-untranslated region (5’-UTR). These methylation steps are crucial for the formation of the type-I cap, crucial during the synthesis of structural and non-structural proteins using the host’s translational machinery [57, 58].
Jung et al. [59] described the development of highly specific RNA aptamers against DENV MTase. Fifteen rounds of SELEX were conducted using an in vitro transcribed RNA library, which has a length of 90 nucleotides. However, the nucleic acid pool obtained from the 13th round showed higher binding affinity towards MTase as compared to the nucleic acid pool from the 15th round. No non-specific binding towards the control Nickel–Nitrilotriacetic Acid (Ni-NTA) agarose beads was observed. Therefore, the nucleic acid pool from the 13th round was used for subsequent cloning and sequencing analysis, yielding 14 candidates with MTase binding and N-7 methylation-inhibition capacities. Only one candidate, aptamer #3, can be truncated while retaining the MTase binding ability.
MulFold platform was used to guide the truncation process by predicting secondary structures after each truncation, resulting in three truncated versions of aptamer #3, which are 59-mer, 45-mer and 33-mer candidates. Only the 59-mer and 45-mer aptamer #3 exhibited N-7 methylation inhibition capacity, despite having much lower Kd as compared to the parent aptamer #3. The loop region of aptamer #3 was predicted to be the interaction site of the aptamer with MTase, as shown by the reduction in the Kd value when targeted mutations were introduced at nine nucleotides in the loop region of the 45-mer aptamer #3. In addition to the binding affinity against DENV-2 that was determined at 28 ± 2.1 nM, the truncated aptamer #3 (45-mer) also binds to the DENV-3 MTase, with a Kd value estimated at 15.6 ± 1.03 nM. However, the truncated aptamer shows no cross-reactivity against JEV MTase and human mRNA cap guanine-N7 methyltransferase (hRNMT). This binding ability can be translated into a notable N-7 methylation inhibition of DENV serotype 3 by the aptamer. However, the truncated aptamer #3 shows no binding to DENV serotypes 1 and 4. The inhibition of N-7 methylation will, in return, halt the type-I cap formation, which thereafter stalls the replication of the virus and promotes the type-I interferon signaling initiation. Aptamer #3 can be a potential therapeutic agent of the Dengue virus.
5’-Untranslated region (5’-UTR)
The untranslated regions (UTR) flank the flavivirus genome at both the 5’ and 3’ ends, consisting of 100 and 400 to 700 nucleotides, respectively [60]. The 5’-UTR is essential for the methylation by DENV MTase, which allows the formation of the type-I cap required for subsequent viral replication [57]. This site is highly conserved across all four DENV serotypes, although some variations have been observed in DENV-3 [61].
Cnossen et al. [62]. demonstrated the isolation of novel aptamer targeting the DENV 5’-UTR using a 55-mer target, which was formed by combining the 5’-UTR with an additional 55 nucleotides required for viral RNA translation. A total of 24 aptamer candidates with varying sizes (41 to 187-mer) were identified after eight SELEX rounds. Alignment analysis revealed the presence of a 5’-GGGGG-3’ motif in most of these aptamers, with candidates of 31 to 40-mer range having this motif. This motif is similar to the 5’-GGGG-3’ motif present in aptamers against the E3 protein of DENV, as reported by Chen et al. [43]. The authors concluded that this motif is vital for recognizing the 5’-UTR and potentially retains this capability. Three shortlisted aptamers, designated A03, B07 and C10, have a loop region following secondary structure prediction by the Nupack platform. These aptamers displayed binding to all DENV serotypes, as demonstrated during the dot blot hybridization assay. In silico analyses also showed that the interaction of the aptamers with the DENV 5’-UTR alters its conformation, suggesting a potential role in inhibiting viral transcription, translation and replication.
Limitations of aptamers as DENV therapeutic agents and strategies to overcome them
While the aptamers have been documented to exhibit therapeutic capacity in inhibiting dengue virus, targeting key viral targets such as E3, methyltransferase and capsid protein, a comprehensive understanding of their robustness remains limited. The lack of studies focusing on their therapeutic potential created a knowledge gap, which then diminishes the possibility of the aptamers-based therapies being translated into preclinical and clinical stages. To date, only two aptamer-based drugs for age-related macular degeneration (AMD), Macugen (pegaptanib) and avacincaptad pegol (Izervay), have been approved for safe usage [63, 64].
One of the key challenges are their moderate vivo stability and rapid clearance. Aptamers are susceptible to degradation by nucleases in biological fluids and are quickly eliminated by renal filtration due to their small size. These factors significantly reduce their circulation half-life and therapeutic efficacy [22, 26, 65]. To overcome this, chemical modifications such as 2′-fluoro, 2′-O-methyl, 3’-inverted dT residue, amino acid-like side chains or locked nucleic acids (LNA) can enhance nuclease resistance [66]. Conjugation with larger molecules like polyethylene glycol (PEG) or liposomes can also help extend their systemic circulation [37, 67, 68]. These refinements, however, are known to be expensive and intricate. In addition, the delivery of anti-DENV aptamers targeting the infected cells is challenging due to the possibility of multicell infections.
Another major hurdle is the efficient delivery of aptamers to target tissues, particularly in reaching intracellular viral components. Systemically administered aptamers may exhibit poor tissue penetration or non-specific distribution. To address this, nanocarrier-based systems such as liposomes, dendrimers, or gold nanoparticles have been explored to improve stability and delivery efficiency [27, 28]. Moreover, the usage of chimeric aptamers, aptameric constructs comprising several individual aptamers that coalesce to perform an integrated function is a viable delivery strategy. For example, a bivalent aptamer that consists of one aptamer targeting the receptor protein and another aptamer for therapeutic purpose can also be used for delivery [66]. Additionally, attaching aptamers to cell-penetrating peptides or targeting ligands may enhance intracellular uptake [42].
Next, a superficial understanding of the pharmacokinetics of these aptamers impedes the development of highly effective and biocompatible aptamer-based therapeutics [69, 70]. Accurate insights into the uptake, distribution, and elimination of aptamers from the body are needed to overcome this issue. Ding et al., 2023 have carried out the first-in-human pharmacokinetics study of sgc8, which is an aptamer targeting protein tyrosine kinase 7, by tagging it with gallium-68 (68Ga). The study has shed some light on the biosafety and pharmacokinetic aspect of sgc8 in the human body [71].
DENV, a virus within the diverse flavivirus family, contributes to the challenge of developing high-affinity DENV-aptamers, which may lead to unwanted cross-reactivity with other non-target proteins [73,72,74]. The time-consuming in vitro assays to validate the specificity of the generated aptamers further exacerbate this issue. Despite this, the availability of in silico selection of aptamers permits the specificity testing to be executed at a faster rate, regardless of the need for substantial computational power [75].
Although aptamers are highly specific, there remains a risk of off-target binding or unintended immune stimulation, especially when perceived as foreign nucleic acids by the host. This issue can be mitigated through careful in vitro and in vivo specificity testing and by incorporating negative selection and careful screening of the candidates during the SELEX process [35]. Chemical backbone modifications may further reduce immunogenicity [37].
Manufacturing and regulatory challenges also pose barriers to the widespread therapeutic use of aptamers. While aptamers are easier to be synthesized than antibodies, ensuring consistent large-scale production and meeting regulatory standards can be demanding. Establishing standardized manufacturing protocols and validated quality control processes is essential, along with continued collaboration with regulatory authorities who are gradually expanding frameworks for oligonucleotide-based therapies [23].
A lack of extensive clinical data, particularly in the context of dengue virus. Most aptamer-based treatments remain in the preclinical stage. Advancing these therapies will require robust in vivo studies and well-designed clinical trials to demonstrate safety and efficacy. Collaborative efforts across academia, industry, and government sectors can help accelerate the clinical translation of aptamer-based dengue therapeutics [29, 33].
Lastly, a huge proportion of anti-DENV research focuses on the development of diagnostics rather than therapeutics, reflecting the funding and technical feasibility for diagnostic-based studies [76]. The development of anti-DENV aptamers as an alternative therapeutic necessitates enormous financial resources to support the extensive validations, ongoing clinical trials and optimized delivery systems vital to ensure high specificity, safety and effectiveness– all result in limited exploration in this specific area.
In summary, while aptamers hold strong potential as therapeutic agents against the Dengue virus, overcoming these limitations through strategic innovations and interdisciplinary cooperation is critical for their future clinical success.