Rebecca Brahde and
Ashlea TraceyIsle of Man
 CHIARA MAZZONE
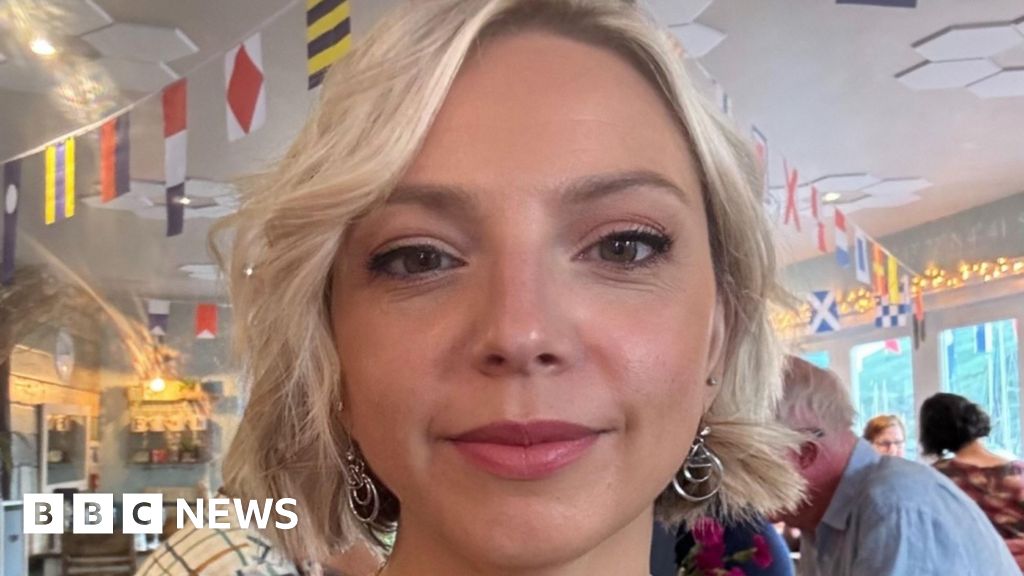
CHIARA MAZZONEA breast cancer survivor has urged others “not to be scared to get checked out”.
Chiara Mazzone had been attending annual…

Rebecca Brahde and
Ashlea TraceyIsle of Man
 CHIARA MAZZONE
CHIARA MAZZONEA breast cancer survivor has urged others “not to be scared to get checked out”.
Chiara Mazzone had been attending annual…
LONG BEACH – Competing in the greater Los Angeles area for the second time in three weeks, the San Diego…
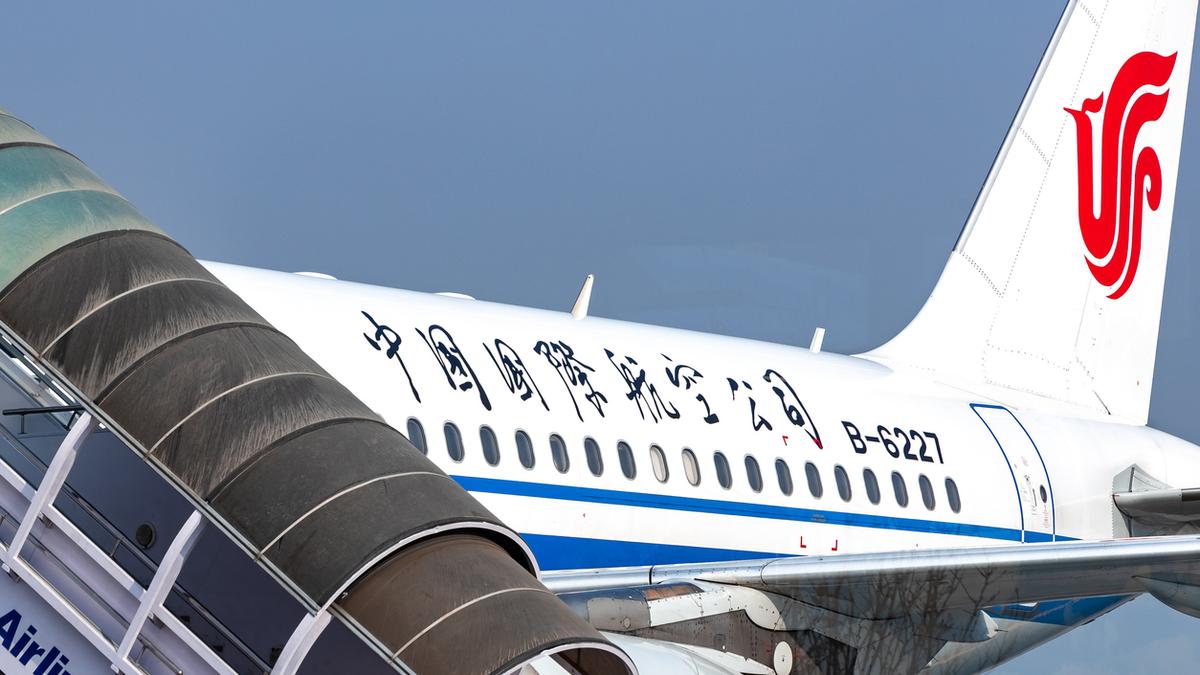

Representative image
| Photo Credit: Getty Images/iStockphoto
A commercial passenger flight operated by Air China was safely diverted to Shanghai on Saturday (October 19, 2025) after a battery stowed in a passenger’s carry-on luggage caught fire, the airline said.
The incident occurred aboard the national carrier’s daily flight from the eastern Chinese city of Hangzhou to Incheon International Airport, near Seoul, South Korea.
“A lithium battery spontaneously ignited in a passenger’s carry-on luggage stored in the overhead bin on flight CA139,” the airline said in a statement on Chinese social media platform Weibo.
“The crew immediately handled the situation according to procedures, and no one was injured,” the statement said.
The plane was diverted for an unscheduled landing at Shanghai Pudong International Airport “to ensure flight safety”, it added.
Bright flames were seen coming from an overhead storage compartment in an image taken by a passenger and published by state-affiliated domestic media outlet Jimu News.
There was black smoke in the cabin, the image showed, as at least one passenger was seen trying to extinguish the blaze.
Data from tracking website Flightradar24 showed that the flight took off from Hangzhou at 9:47 a.m. local time.
It made a complete turn over the sea roughly equidistant from the eastern Chinese coast and Japan’s southern island of Kyushu, landing in Shanghai shortly after 11am local time.
Published – October 19, 2025 11:47 am IST
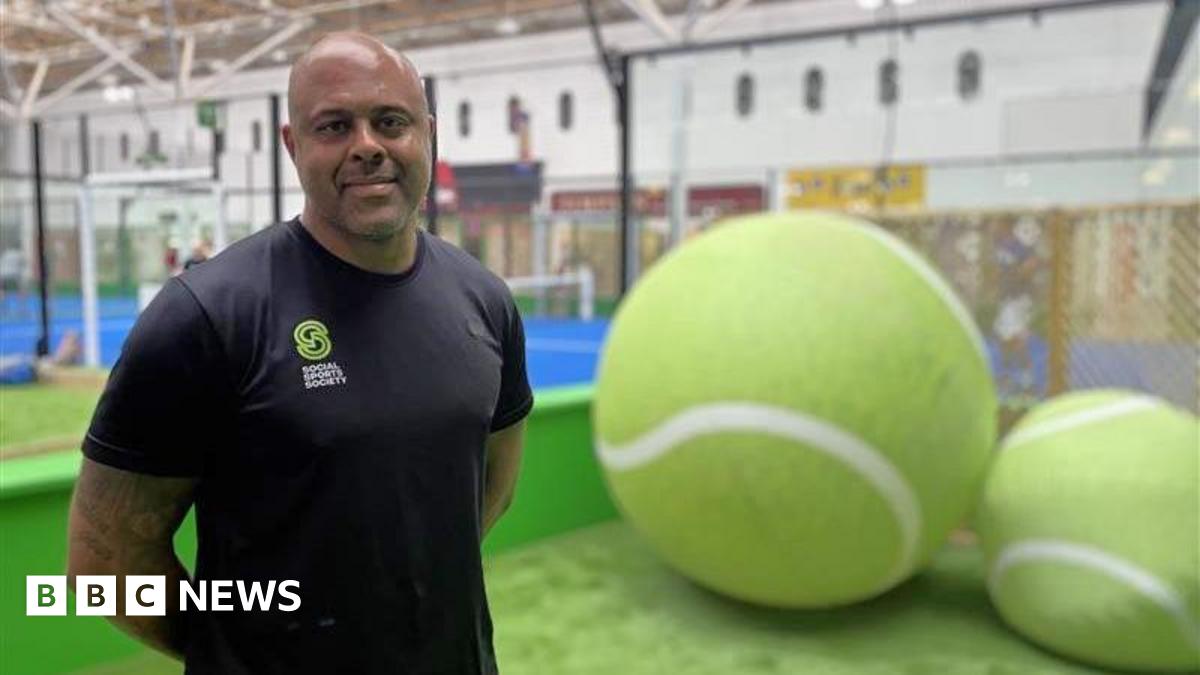

One of the UK’s largest indoor padel centres has opened in place of a former market in Derby city centre.
The centre, which features 10 indoor courts, has regenerated part of the former Eagle Market space, with the remaining area set to become a…
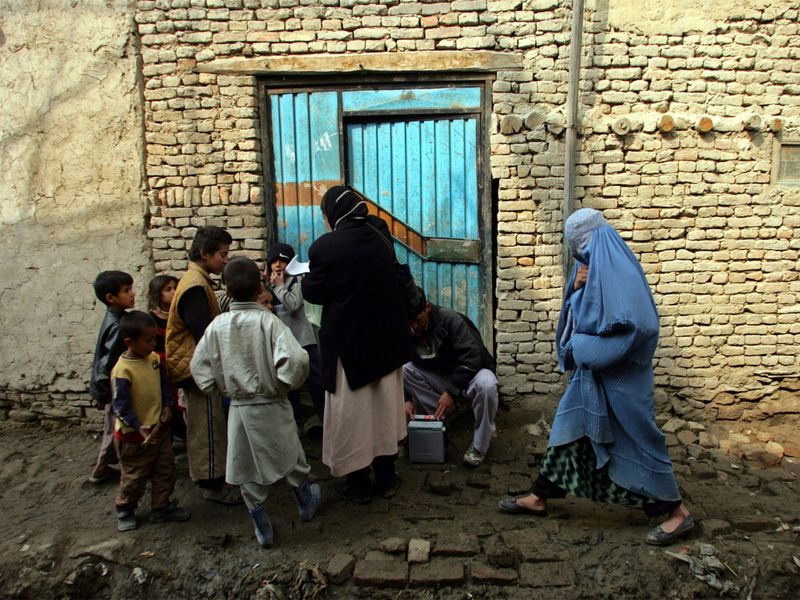

Kabul [Afghanistan], October 19 (ANI): Afghanistan has launched the polio vaccination drive for the country’s eastern provinces, as reported by Khaama News on Saturday.
As per Khaama News, the polio campaign began in Afghanistan’s eastern…

New Delhi: India’s much-touted “independent foreign policy” has once again been exposed as hollow rhetoric, as New Delhi has finally succumbed to U.S. pressure and halved its oil imports from Russia — a move that underscores its…

Can artificial sweeteners actually be harmful to health? And can consuming them on the regular affect our memory and thinking skills? Although a new link found between sugar alternatives and brain aging requires more study, it may warrant some…
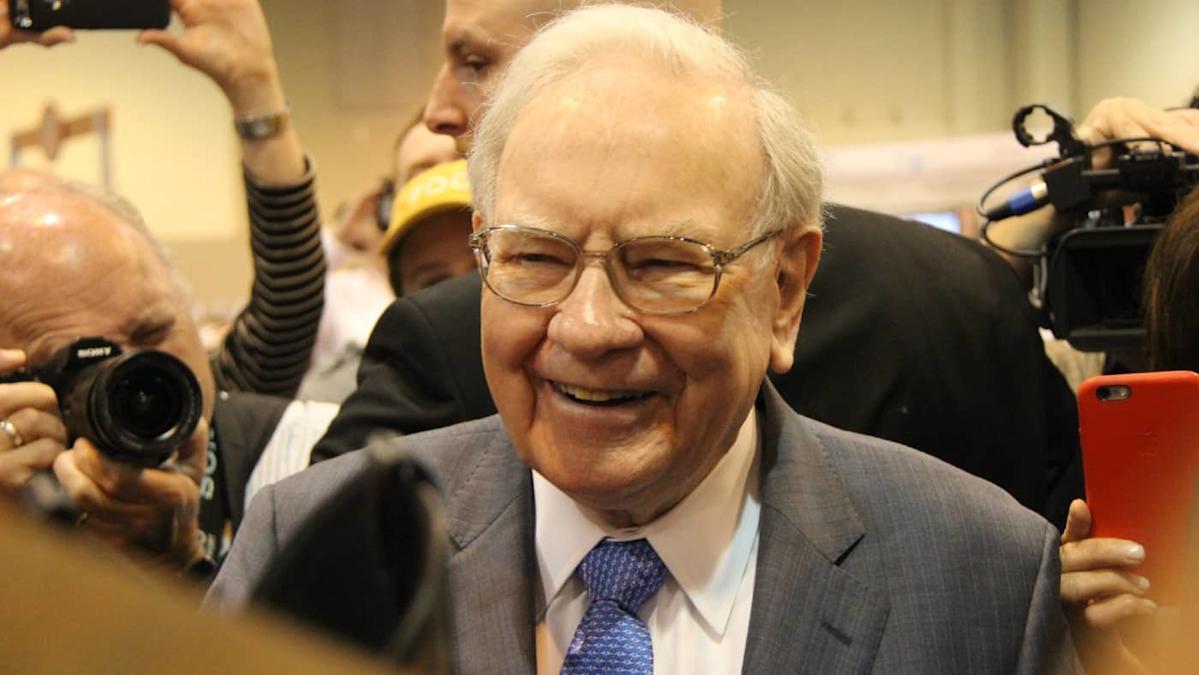

With US stock prices reaching new record highs, the Warren Buffett Indicator has reached an alarming 218%. As a reminder, when it rises above 160% it’s historically suggested that US stocks are significantly overvalued.
This isn’t the only signal hinting that valuations might be getting stretched in 2025. And yet, despite rising concerns, the ’Oracle of Omaha’ and his team are still buying some US shares.
So what stocks is the billionaire investor buying? And should investors follow in his footsteps?
With a reputation for being a value-oriented investor, the fact that Buffett’s buying at a time when valuations are high seems strange, on the surface. But digging deeper, he’s actually still executing the same tried and tested strategy of prioritising value at a fair price.
Two of his recent investments in Nucor (NYSE:NUE) and UnitedHealth Group (NYSE:UNH) seem to demonstrate this perfectly.
So what do these businesses do? And why is Buffett’s team buying them now?
Nucor is a US steel producer. In fact, it’s one of the largest in the country, operating an expansive network of furnaces that recycle scrap metal into quality steel. This unique approach drastically reduces the cost of manufacturing, giving the group a notable competitive edge over its US rivals – something Buffett loves to see.
With the US imposing a 50% tariff on imported steel, Nucor now looks far more attractive to steel consumers. And when combined with surging steel demand courtesy of artificial intelligence (AI) infrastructure and national electrification spending, the business looks like it could be a new champion within the supply chain of countless US-based businesses.
Combining this with an undemanding forward price-to-earnings ratio of 12.2, it’s not so surprising that Buffett‘s taken an interest.
Of course, there are still risks. Even with tariffs, steel demand remains highly cyclical and sensitive to activity within the construction and industrials sectors.
Since higher interest rates often subdue activity within these industries, Nucor’s growth could prove lacklustre, especially if inflation continues to prove sticky. And if AI infrastructure spending starts to slow, the firm could lose a significant tailwind that’s currently driving it forward.
UnitedHealth’s also another seemingly cyclical strategy the billionaire’s pursuing. The firm’s the largest health insurance provider in the US. And with an ageing population, demand for its services is suspected to steadily trend upward.